Medical Terminology Automated Lookup with Pre-Annotation for Healthcare Projects
Generative AI Lab supports integration with Medical Terminologies as a core platform capability, enabling standardized clinical annotation using widely adopted coding systems such as ICD-10, LOINC, CPT, SNOMED CT, RxNorm, and MeSH.
This feature supports both:
- Automated code resolution during pre-annotation
- Manual terminology lookup during annotation and review
Together, these capabilities improve consistency, interoperability, and clinical data normalization across healthcare projects.
Medical Terminologies are integrated into the Hub, Label Configuration, and Cluster Management workflows, allowing teams to discover, download, configure, and deploy terminology resources as shared assets across multiple projects.
By combining automated code resolution with manual lookup capabilities, users can ensure that entity annotations carry standardized medical codes throughout the annotation workflow, from pre-annotation through human review and export.
Discovering and Managing Medical Terminologies in the Hub
Medical Terminologies are available as a dedicated asset type in the Hub, where users can browse available terminologies, understand their coverage, and manage downloads.
A dedicated Medical Terminologies management page provides centralized access to:
- Browse available terminology assets (ICD-10, LOINC, CPT, SNOMED CT, RxNorm, MeSH)
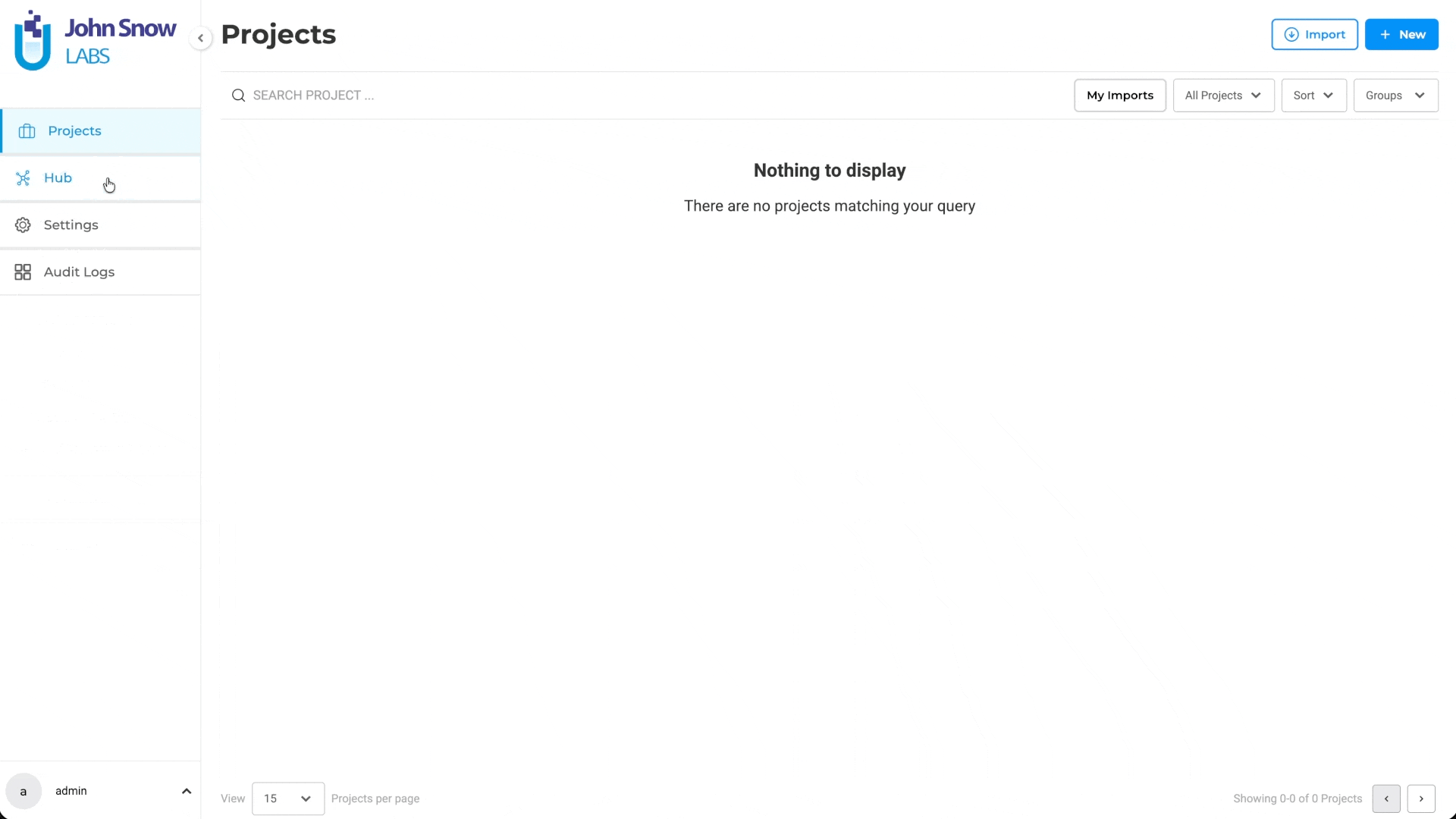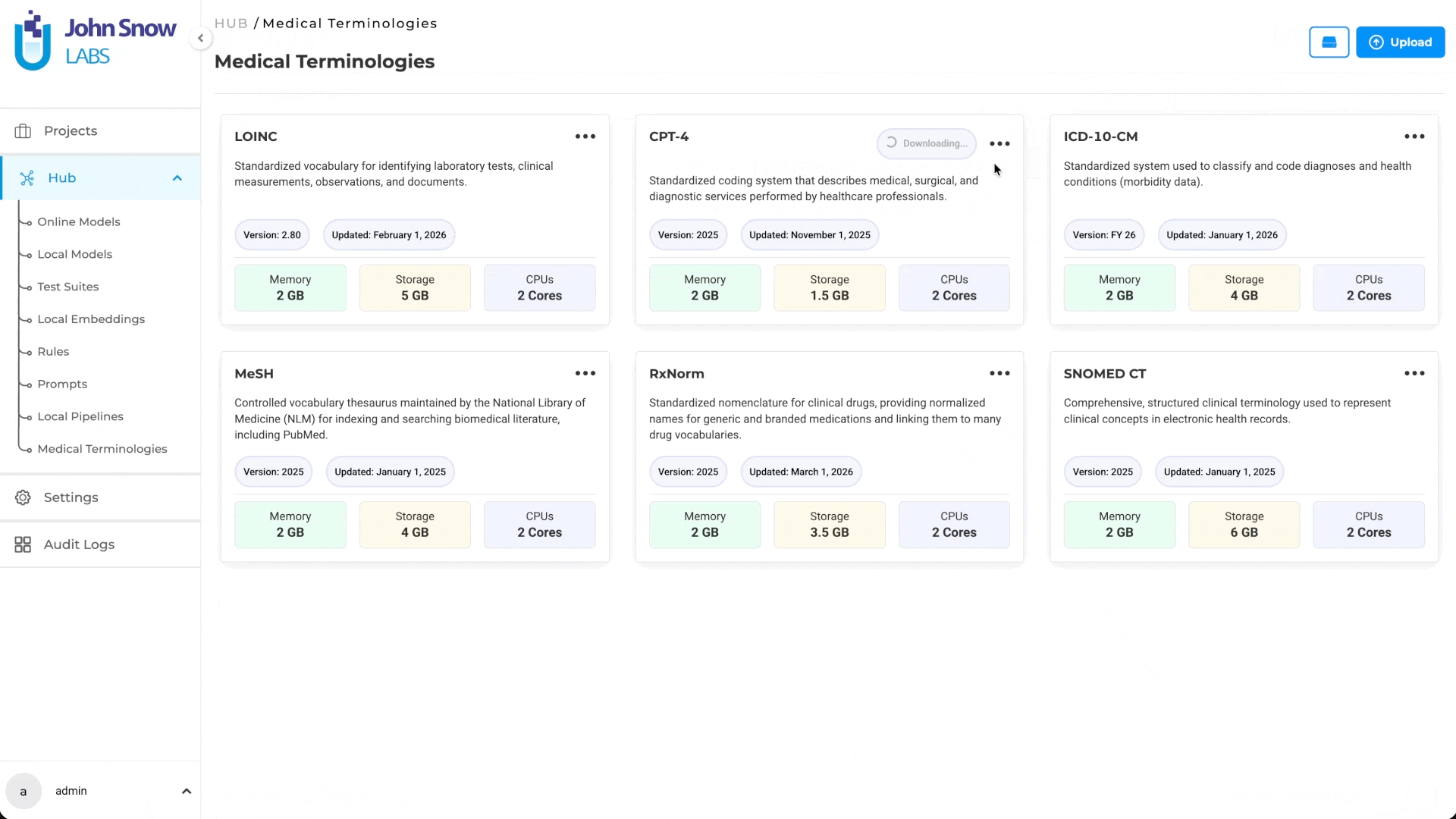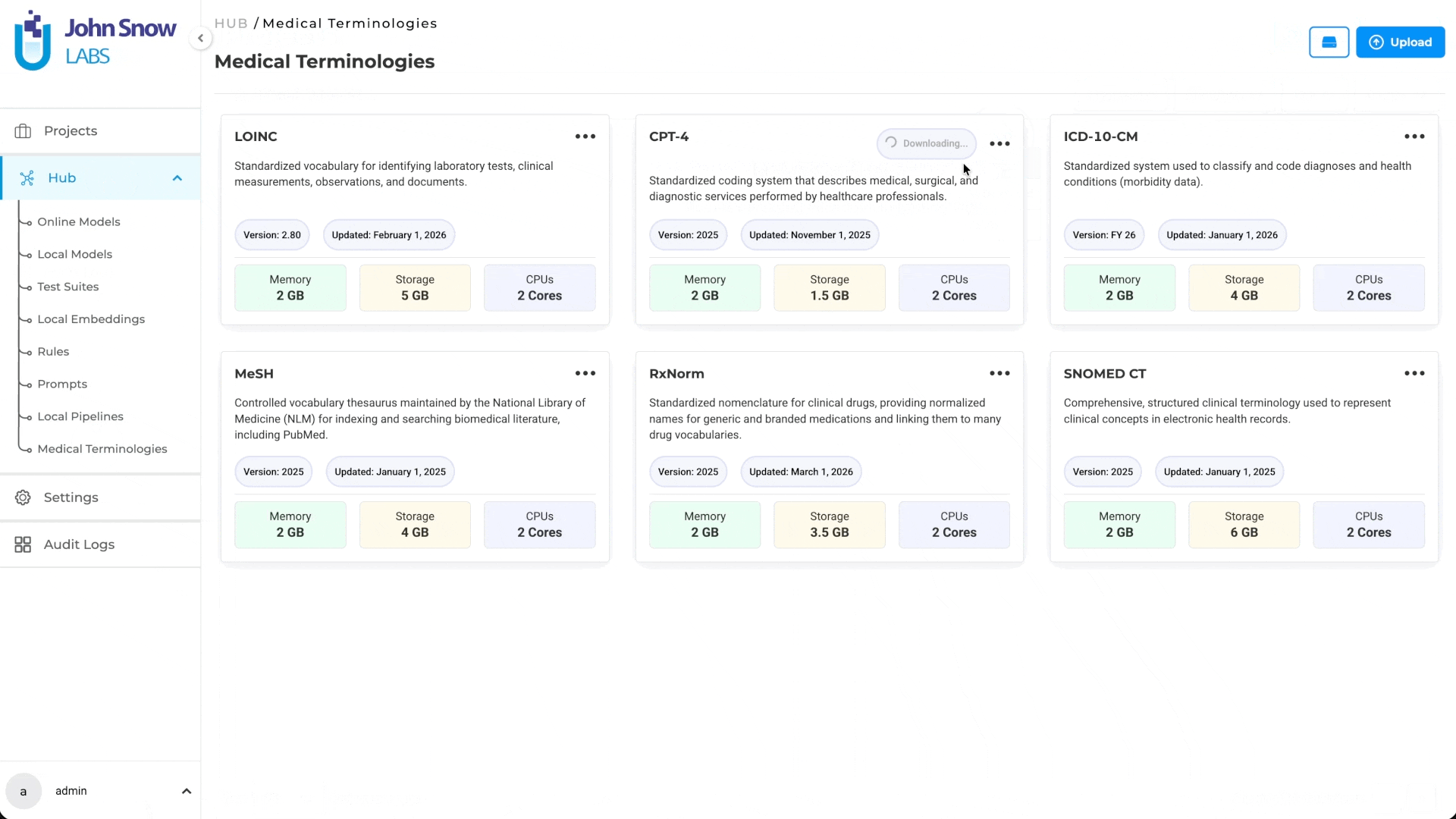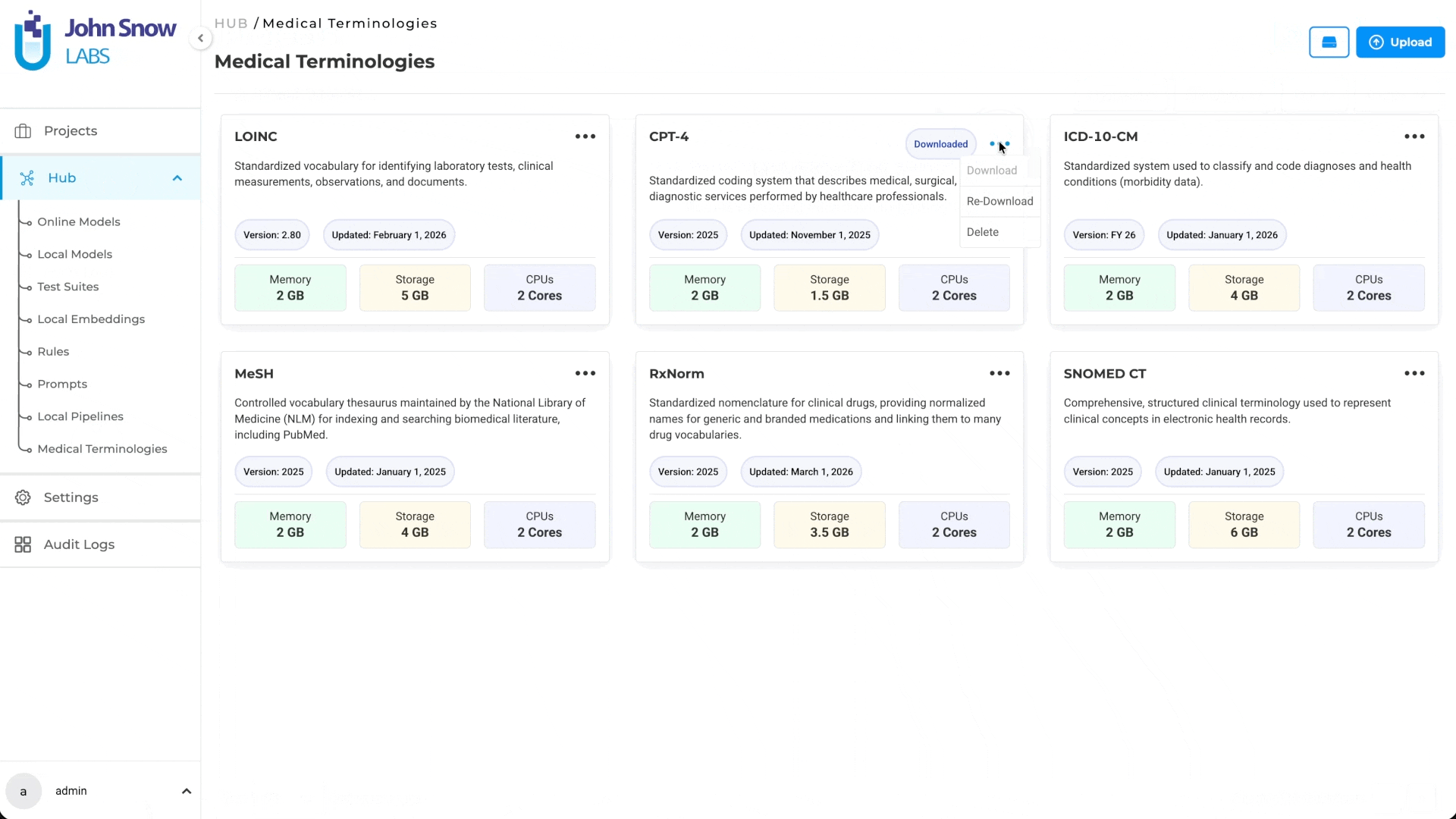
- Download terminologies for use across projects
- Re-download terminologies if needed
- Delete terminologies that are no longer required
- View deployment status (Active indicator appears when a terminology is deployed)
Once downloaded, a terminology becomes a shared resource available across multiple projects. Teams working on different annotation initiatives can draw from the same terminology without redundant setup.
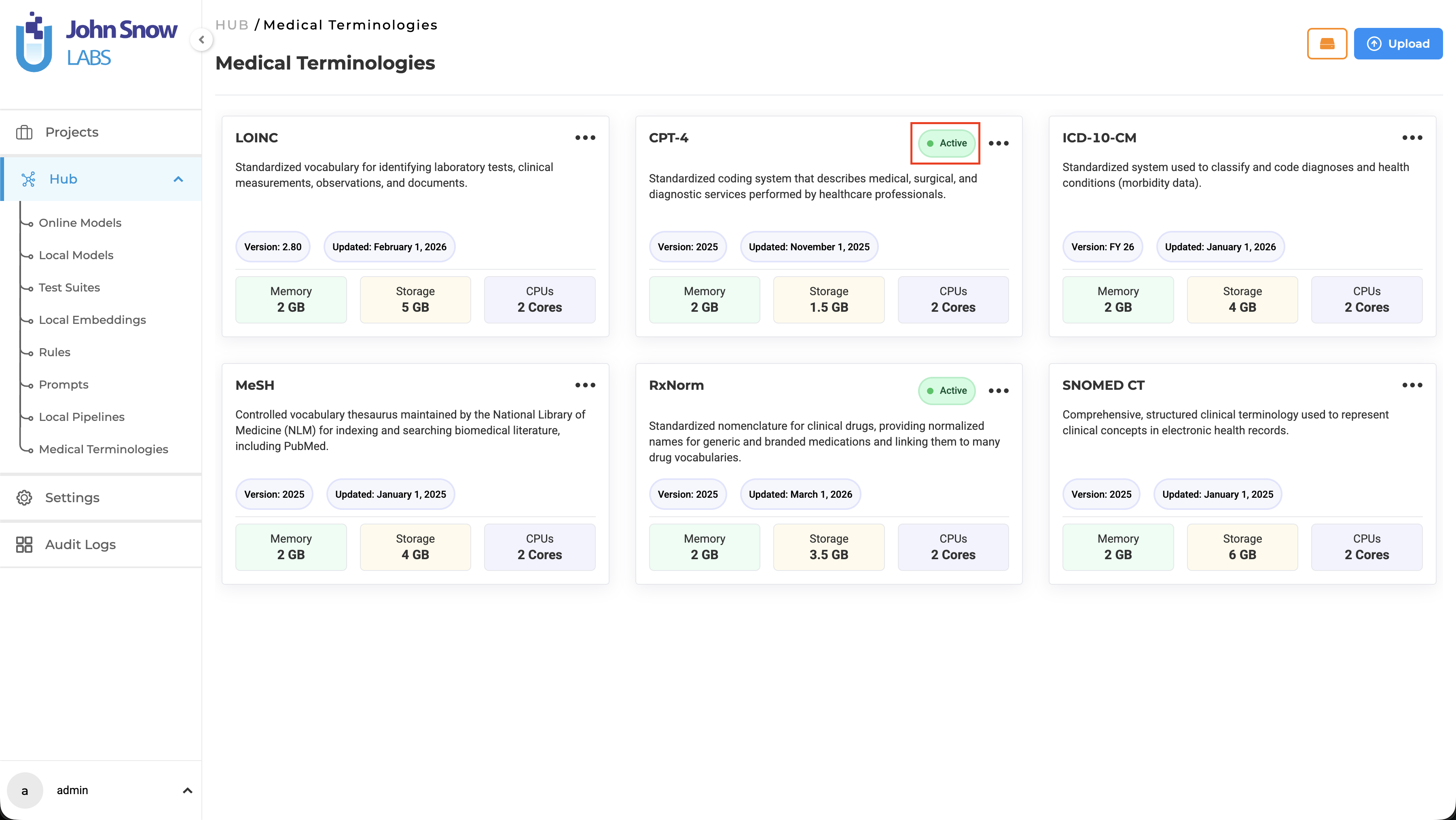
Note: Downloading Medical Terminologies requires a valid license with the appropriate healthcare scope. Downloaded terminologies are available immediately for project configuration without additional infrastructure steps.
Configuring Medical Terminologies in Projects
The integration point between Medical Terminologies and annotation work is the Label Configuration interface. When setting up a project, teams can link specific terminologies to specific labels through the redesigned Customize Label Configuration page.
Steps to Configure Medical Terminologies:
- Create a new project or navigate to an existing project
- Go to Setup → Configuration → Customize Label
- Enable the Medical Terminologies toggle switch
- Map specific terminologies to labels (e.g., ICD-10 to diagnosis labels, LOINC to lab result labels, CPT to procedure labels)
- Save the configuration
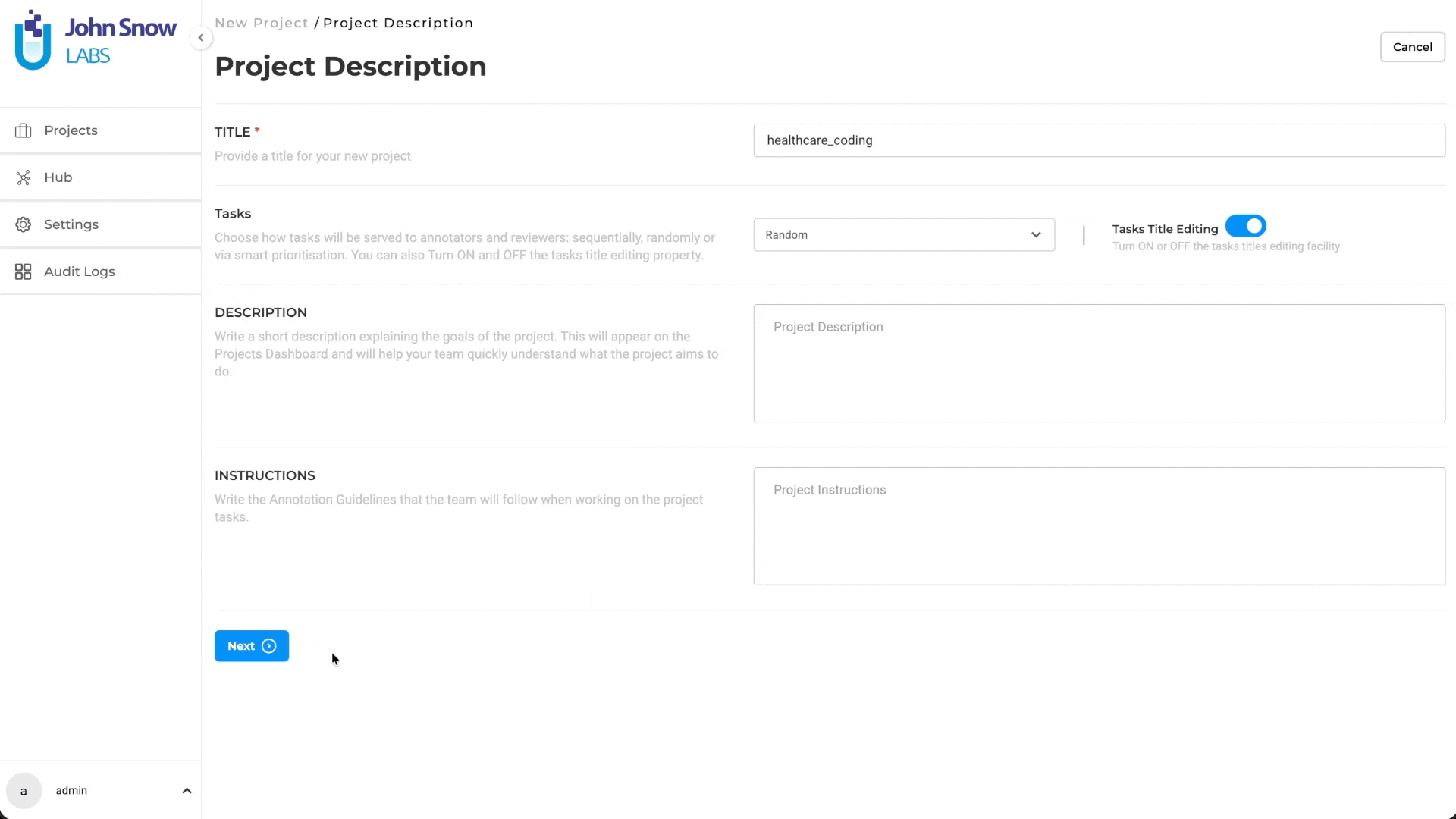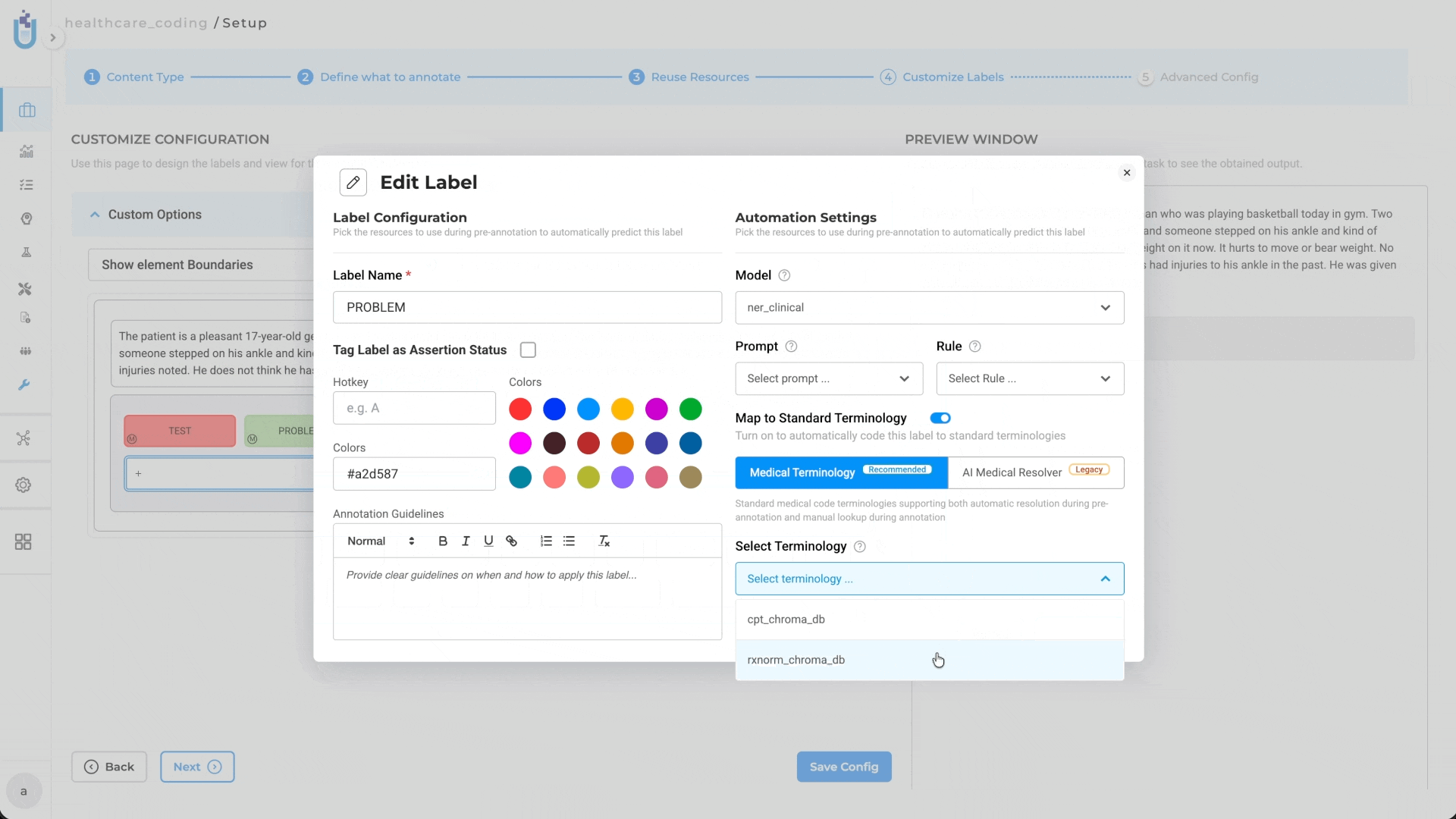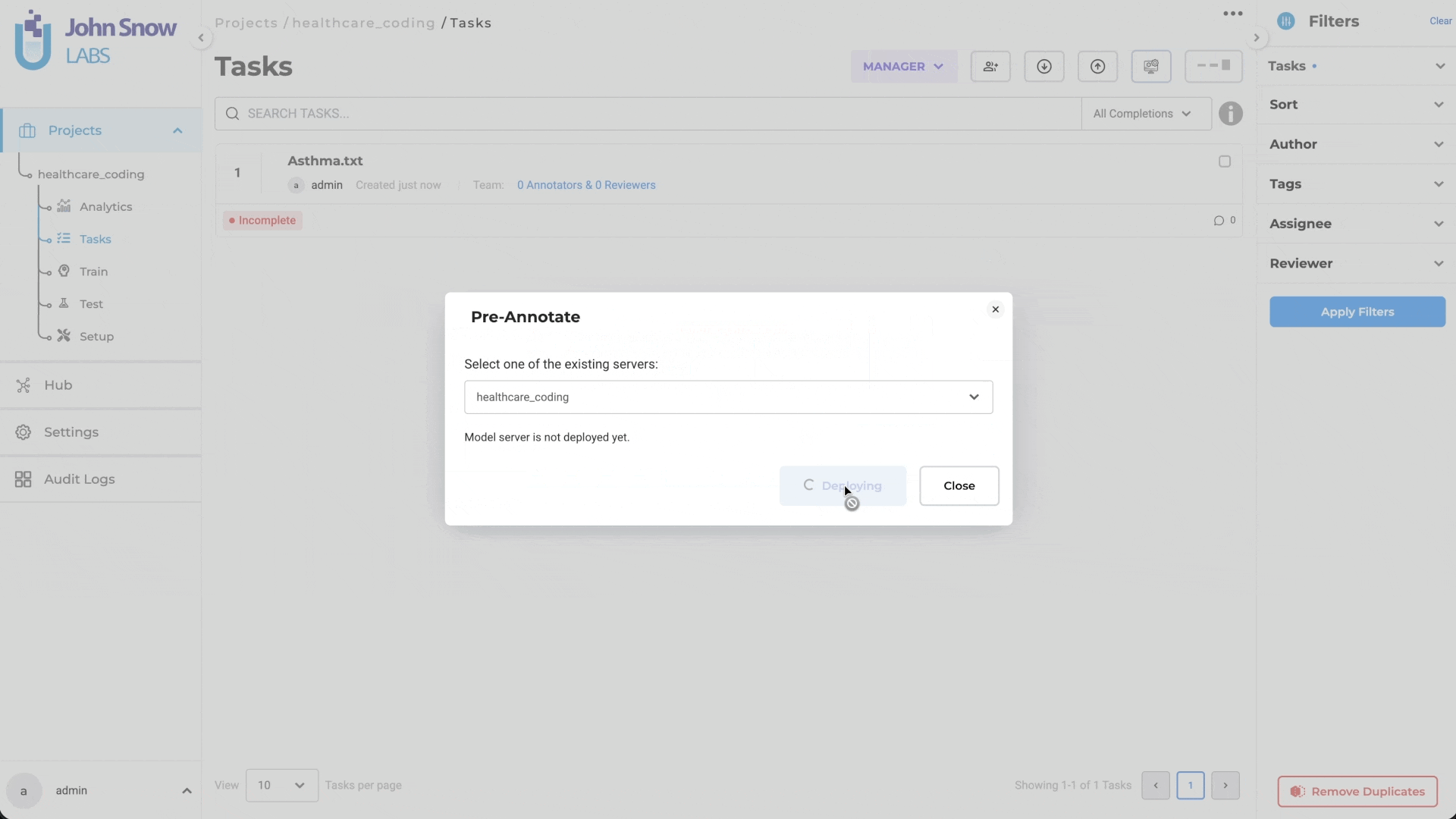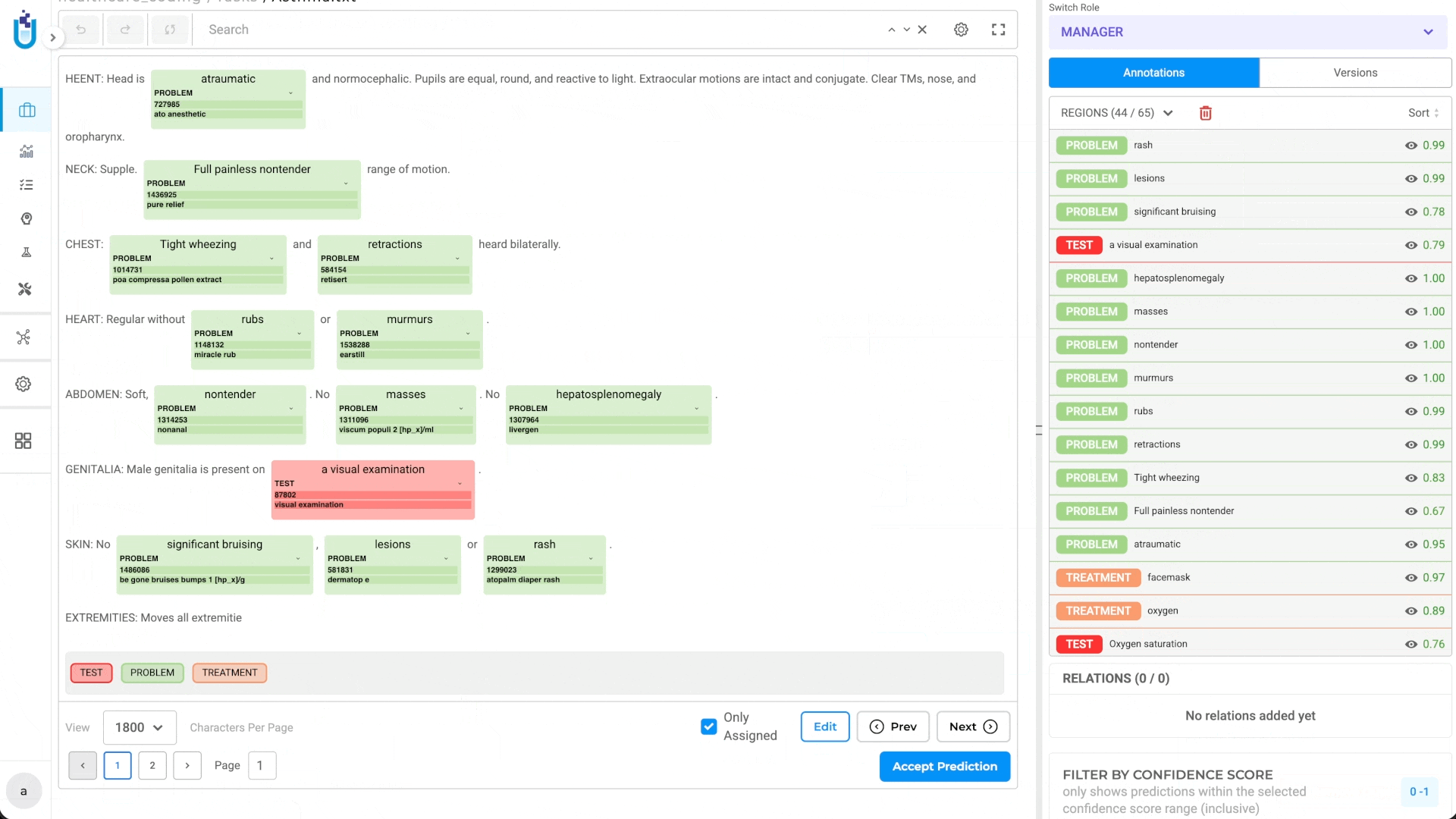
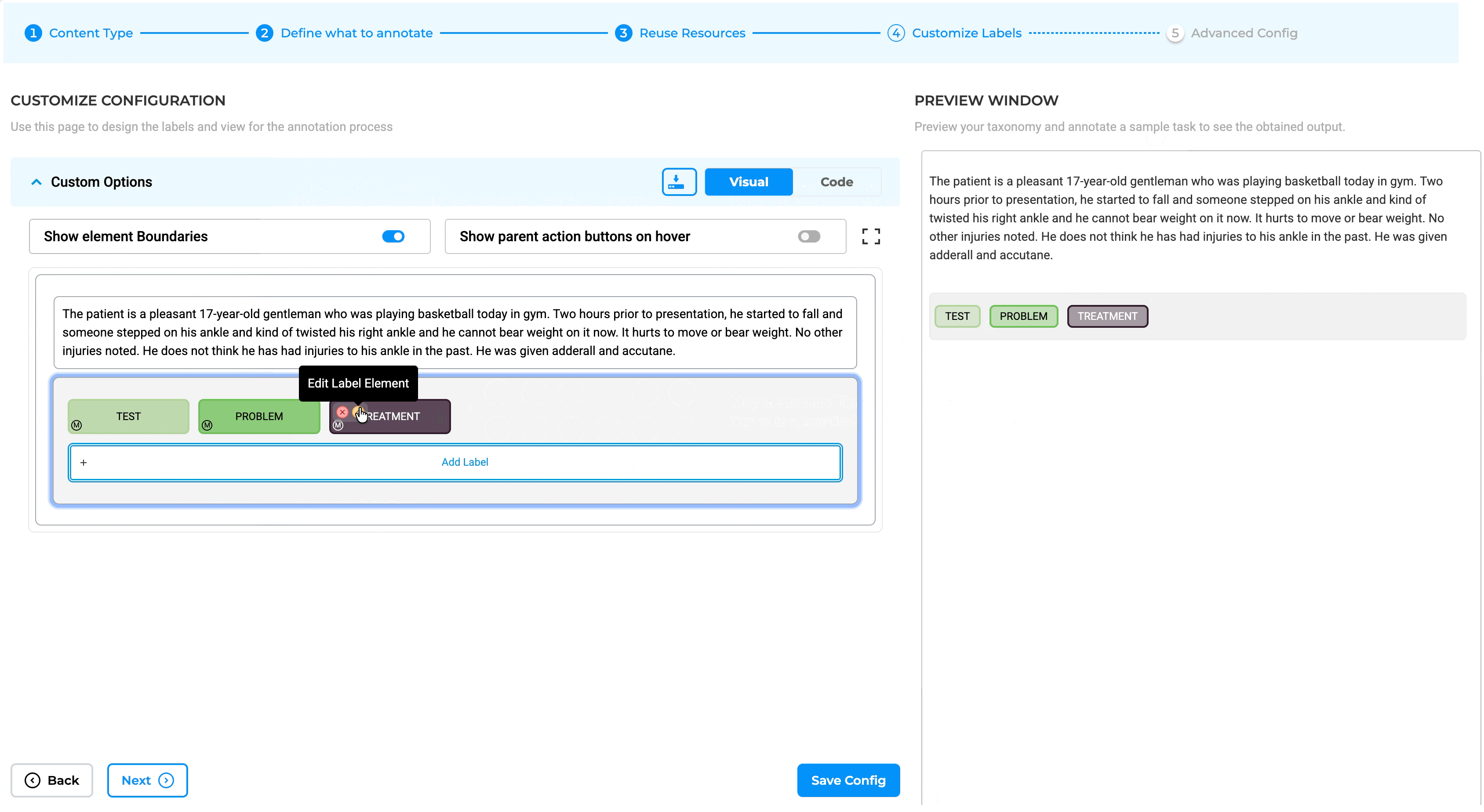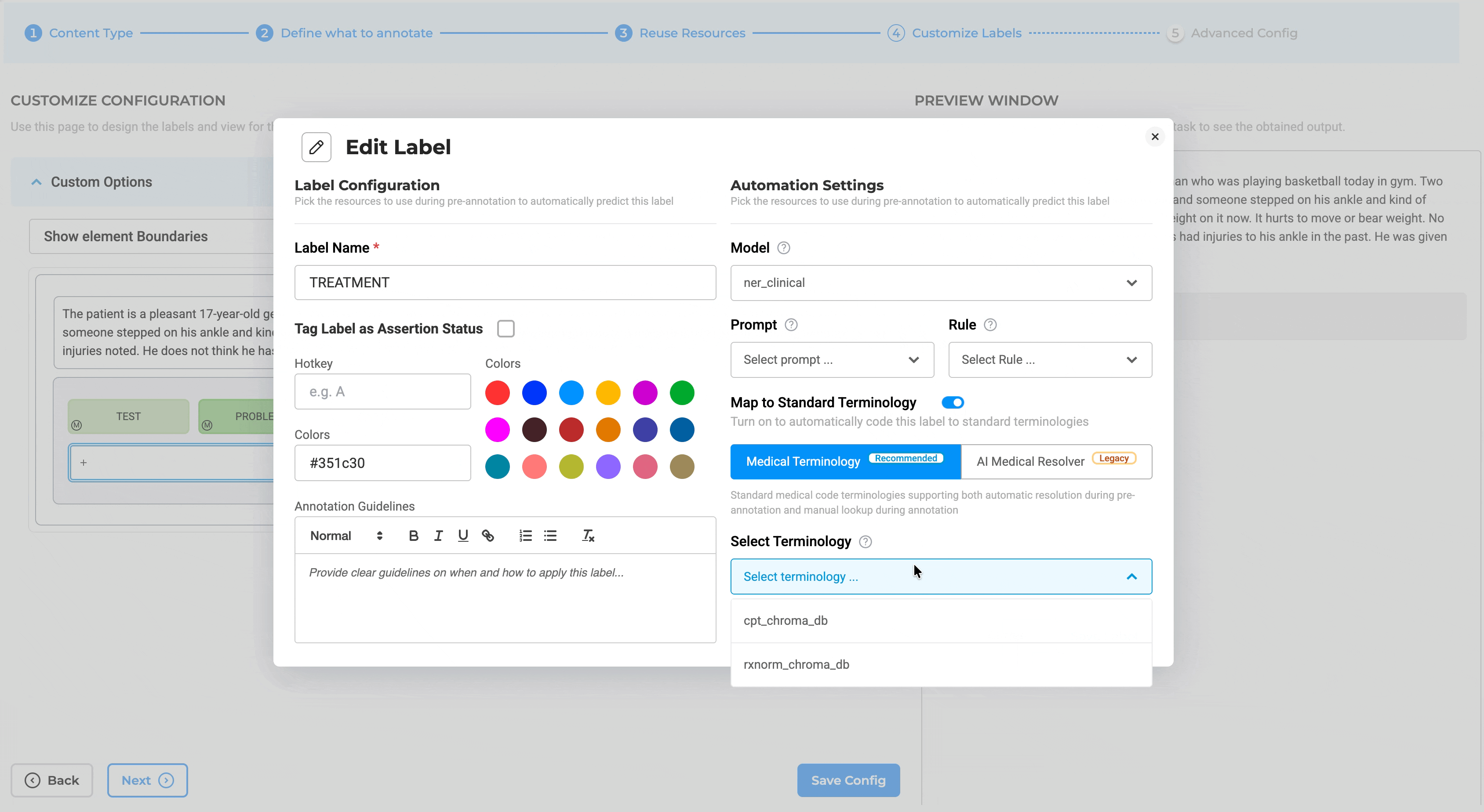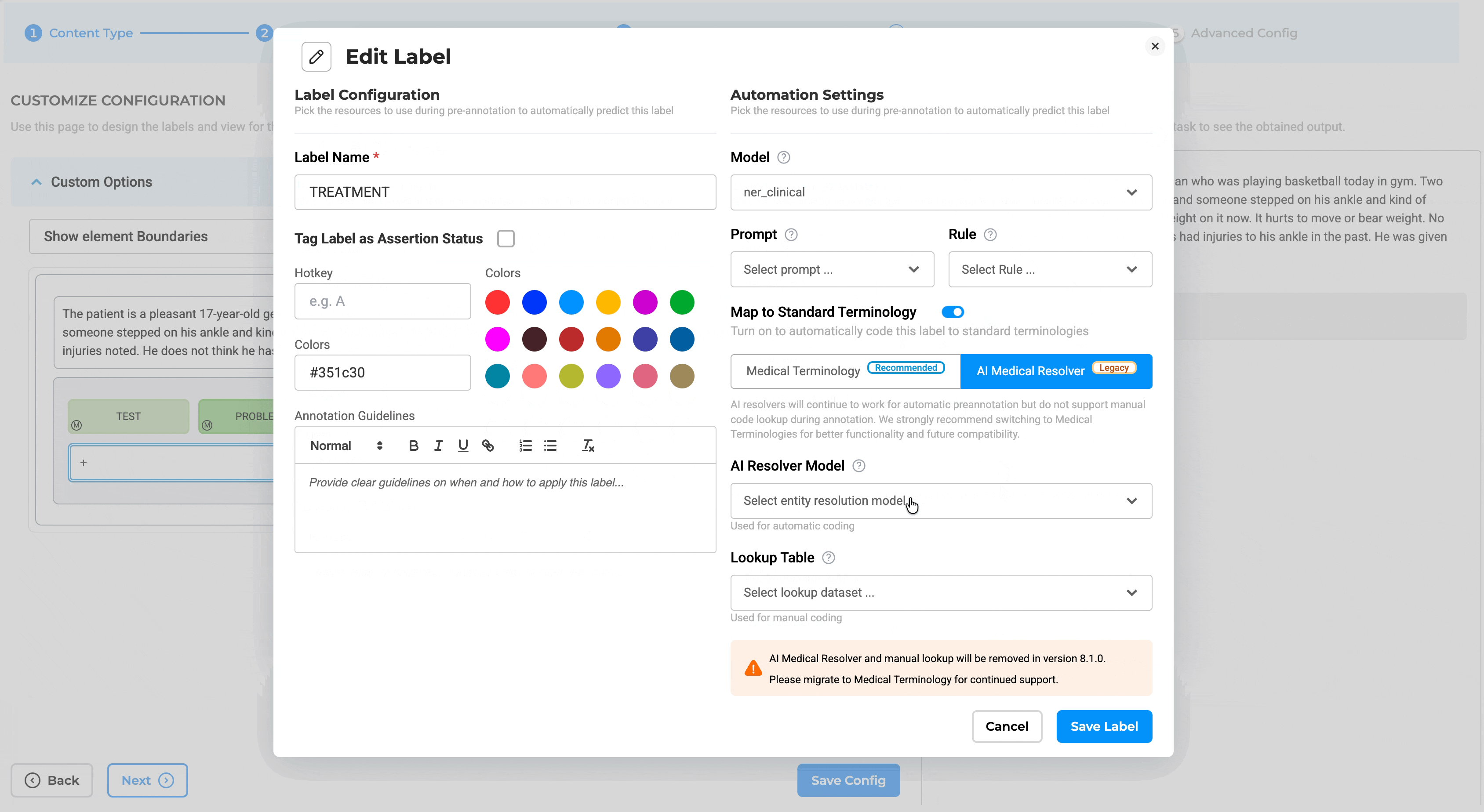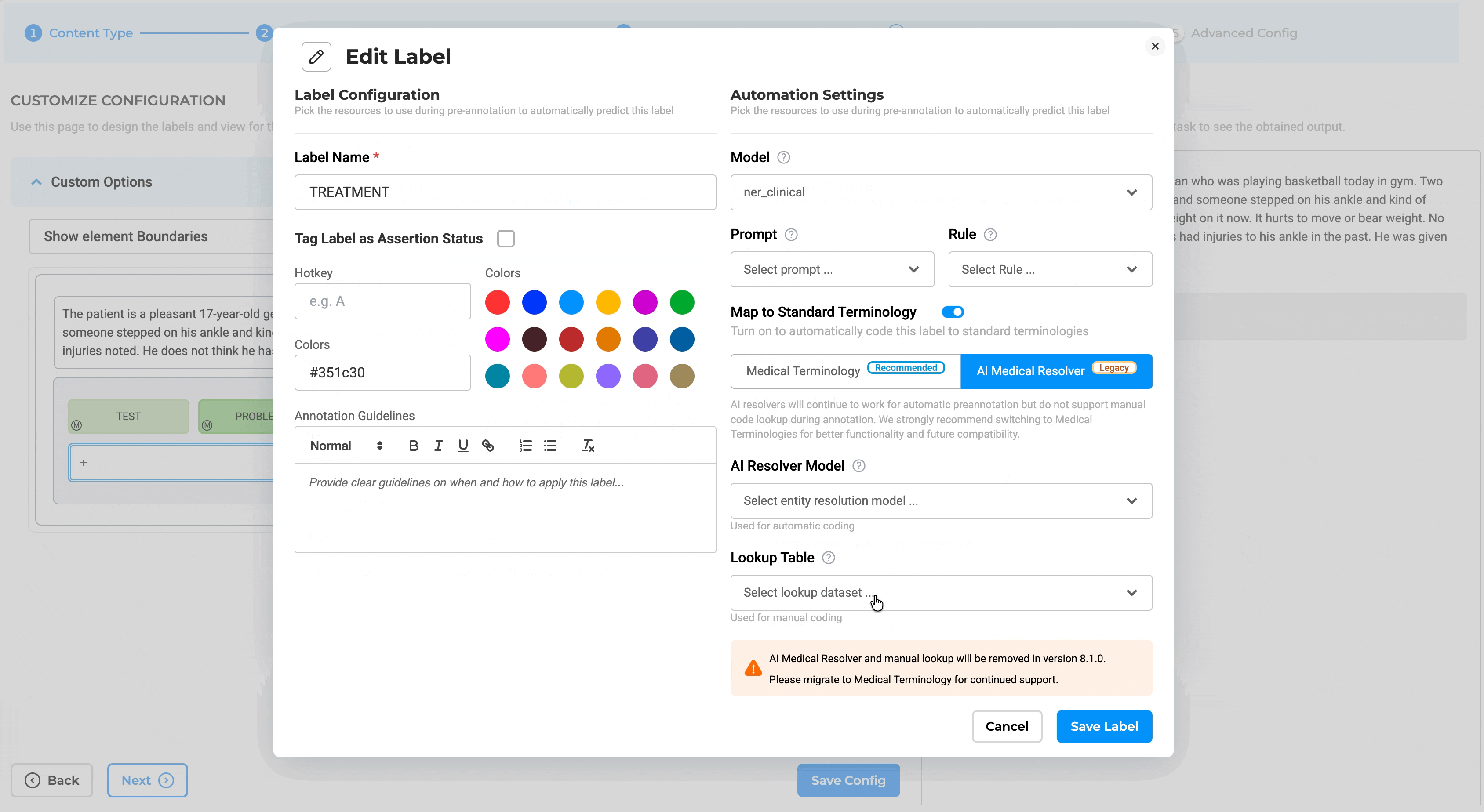
Label-Level Mapping: Projects can link one or more terminologies through Customize Label Configuration, with label-level mapping supporting targeted terminology usage. For example:
- Map ICD-10 to diagnosis labels
- Map LOINC to laboratory observation labels
- Map CPT to procedure labels
- Map RxNorm to medication labels
This label-level association determines how code resolution happens during both automated pre-annotation and manual annotation workflows.
Note: Medical Terminologies cannot be used together with legacy AI resolver models. The system enforces this validation and displays a message if conflicting resolver types are configured.
Deploying Medical Terminology Servers
Once Medical Terminologies are configured in a project, they need to be deployed to enable both automated code resolution and manual lookup functionality.
Deployment Options:
Option 1: Deploy with Pre-Annotation Server
- Navigate to the Task page
- Click the Pre-Annotate button
- Select Create Server from the dropdown
- Click Deploy
- Both the Medical Terminology Server and Pre-Annotation Server are deployed together
Option 2: Deploy Standalone Terminology Server
- Use the Deploy Medical Terminology Server button on the Task page
- Deploys a standalone Medical Terminology Server for manual lookup workflows
- Required when users need manual terminology lookup without pre-annotation
Unified Deployment: When a project links to multiple terminologies, the system automatically deploys them together on a single terminology server to optimize infrastructure usage. This unified deployment reduces resource consumption and simplifies management.
Server Reuse Logic: A project can reuse an existing active terminology server if it requires the exact same set or a subset of the terminologies already deployed on that server. The system automatically identifies compatible servers and reuses them rather than creating redundant deployments.
Note: If the Deploy and Save Configuration button is used while saving configuration that includes Medical Terminologies, only the Pre-Annotation Server is deployed. To enable manual lookup, use the Pre-Annotate button deployment or the dedicated Deploy Medical Terminology Server button.
Automated Code Resolution During Pre-Annotation
Linked Medical Terminologies run automatically during pre-annotation. When the pre-annotation pipeline processes a document and extracts an entity matching a configured label, the system resolves the corresponding medical code as part of the same pass.
How Automated Resolution Works:
- Pre-annotation extracts entities from the text (e.g., “Type 2 diabetes”)
- The system identifies the label associated with the entity (e.g., “Diagnosis”)
- The configured Medical Terminology (e.g., ICD-10) is queried automatically
- The resolved code (e.g., “E11.9”) is attached to the annotation
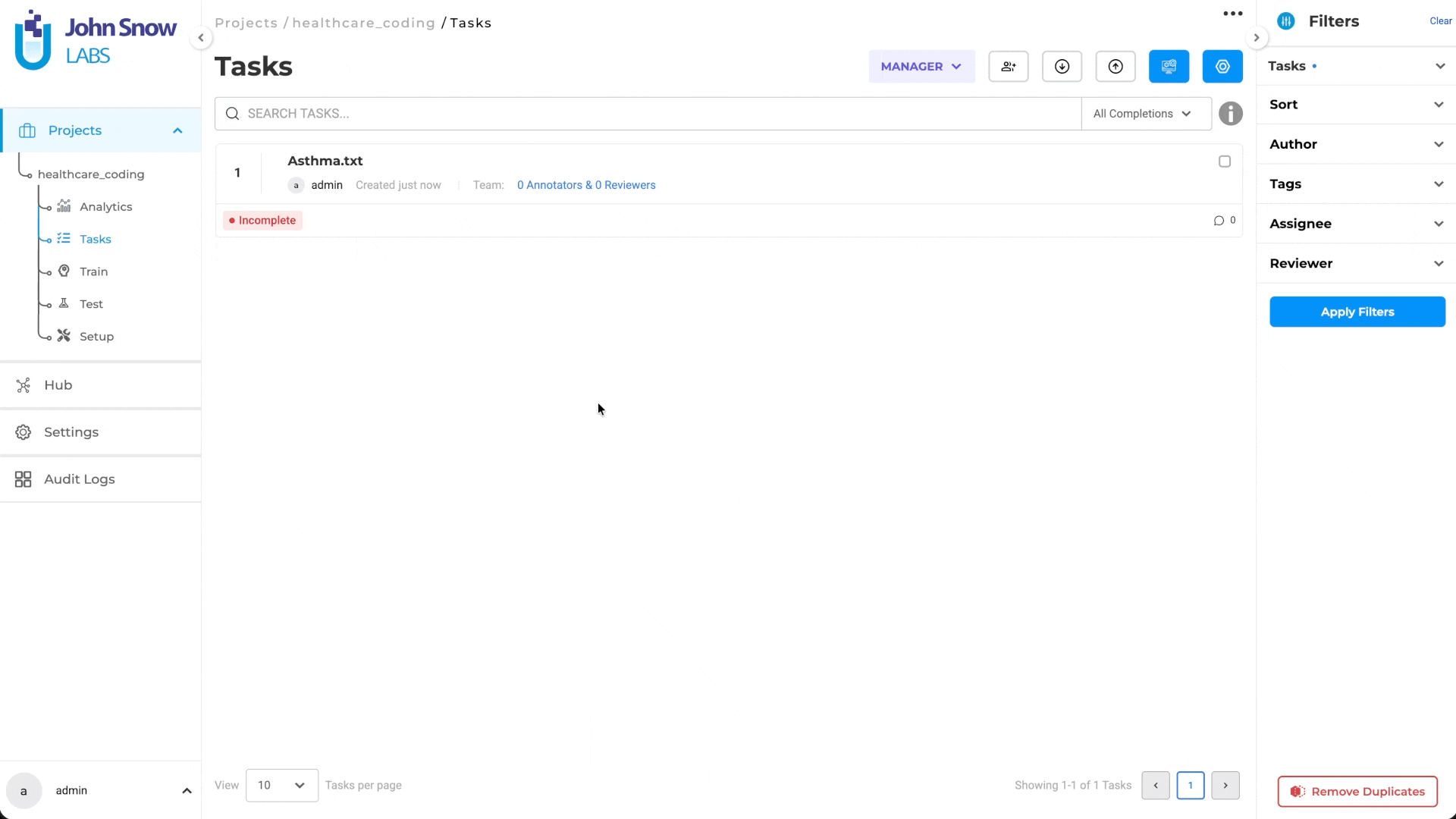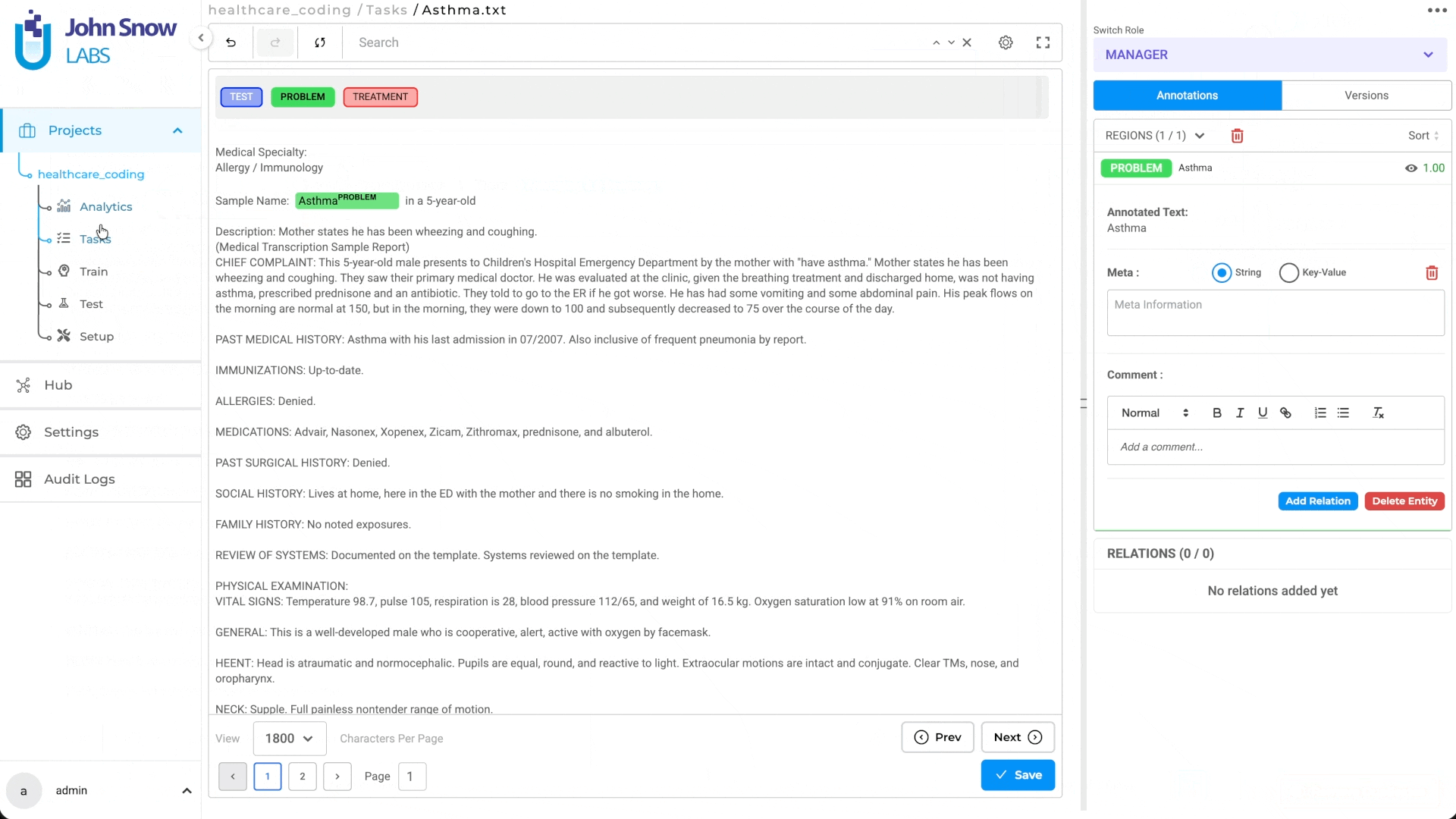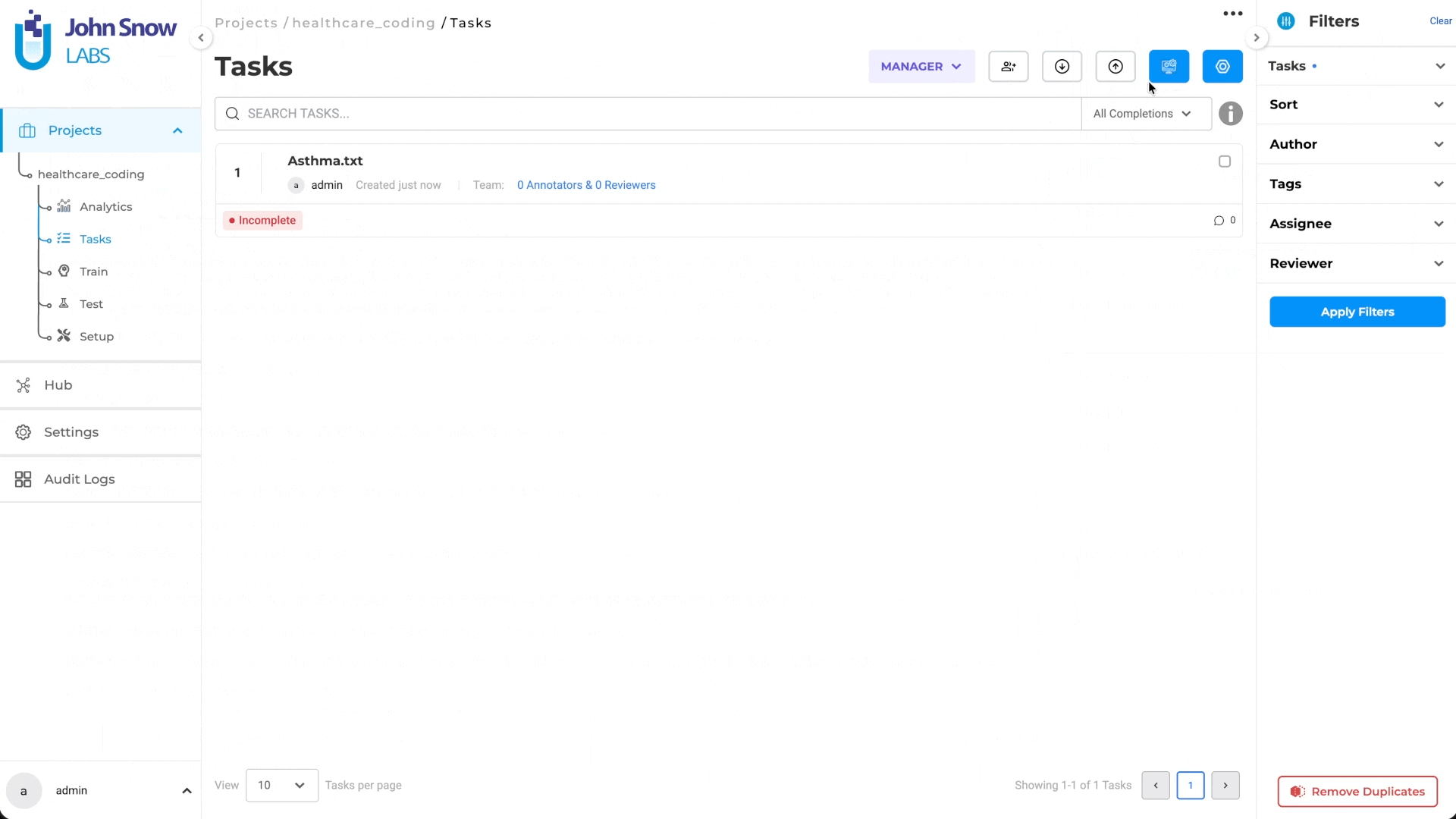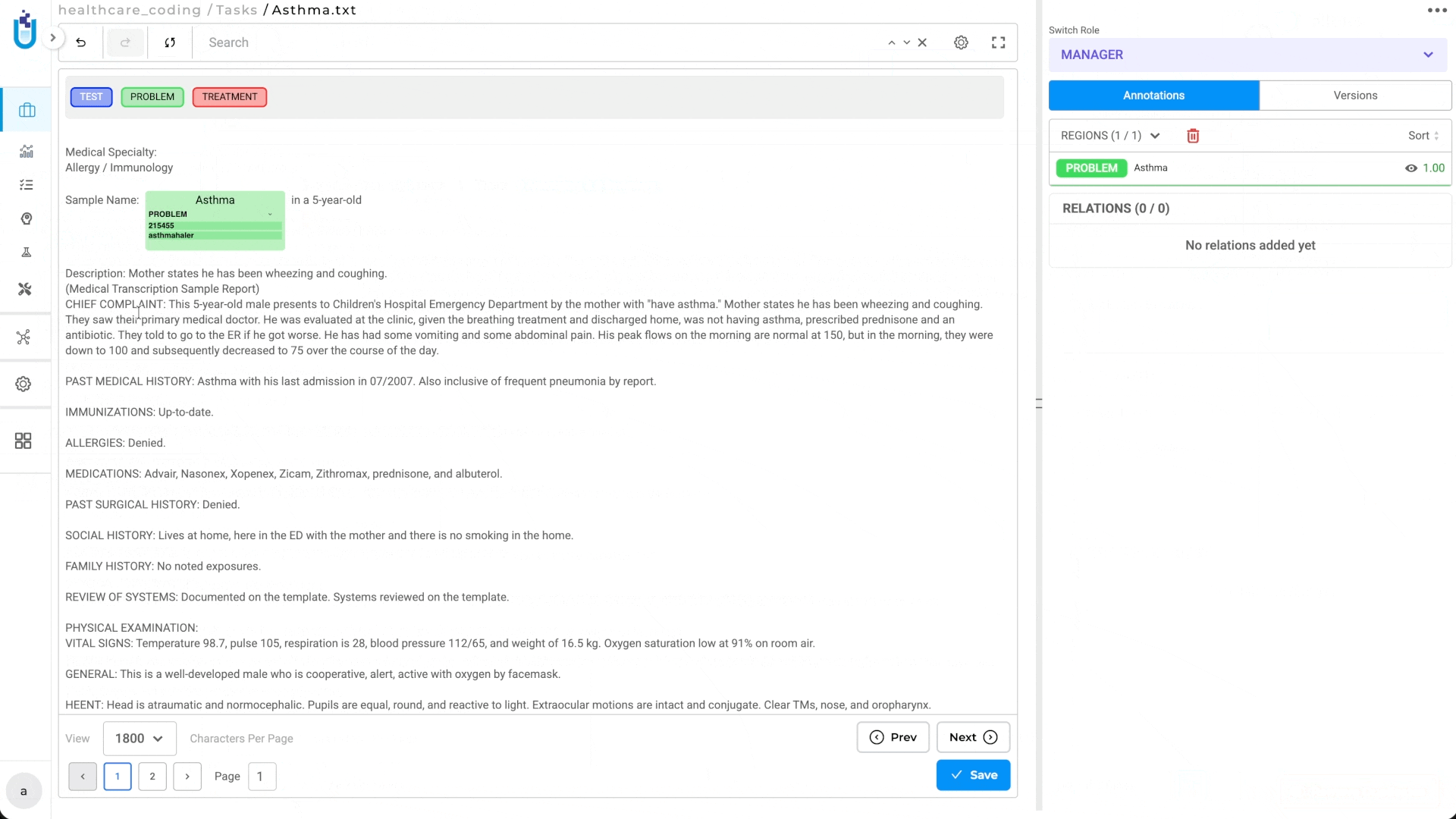
- Annotators reviewing results see the entity span, label, and resolved code together
This automated workflow means annotators see not just the entity and label, but the resolved code—ready to verify or correct, not to look up from scratch.
Benefits of Automated Resolution:
- Reduces manual coding work during annotation
- Ensures consistency in code assignment
- Speeds up annotation throughput
- Maintains the connection between entity, label, and code throughout the workflow
The entity, label, and code travel together through the annotation workflow and into the export, ensuring that downstream systems receive standardized coded data.
Manual Terminology Lookup During Annotation
In addition to automated resolution, Medical Terminologies support manual lookup directly within the annotation interface. Annotators working on entities where automatic resolution didn’t produce a confident result can query the linked terminology interactively.
How Manual Lookup Works:
- Annotator selects an entity span in the document
- Applies the appropriate label
- Opens the terminology lookup interface
- Queries the configured Medical Terminology (e.g., searches for “chest pain” in ICD-10)
- Selects the correct code from the results
- The code is attached to the annotation
When Manual Lookup Is Useful:
- When automated resolution doesn’t produce a result
- When multiple valid codes exist and human judgment is needed
- When reviewing and correcting automated code assignments
- When annotating edge cases or ambiguous entities
Manual lookup keeps annotators within the platform—no need to switch to external coding tools or reference materials. The coded result remains attached to the annotation decision, maintaining workflow integrity.
Resource Configuration and Management
Administrators can configure resource allocation for Medical Terminology Servers through the Infrastructure page. This allows teams to customize CPU and memory settings based on workload requirements.
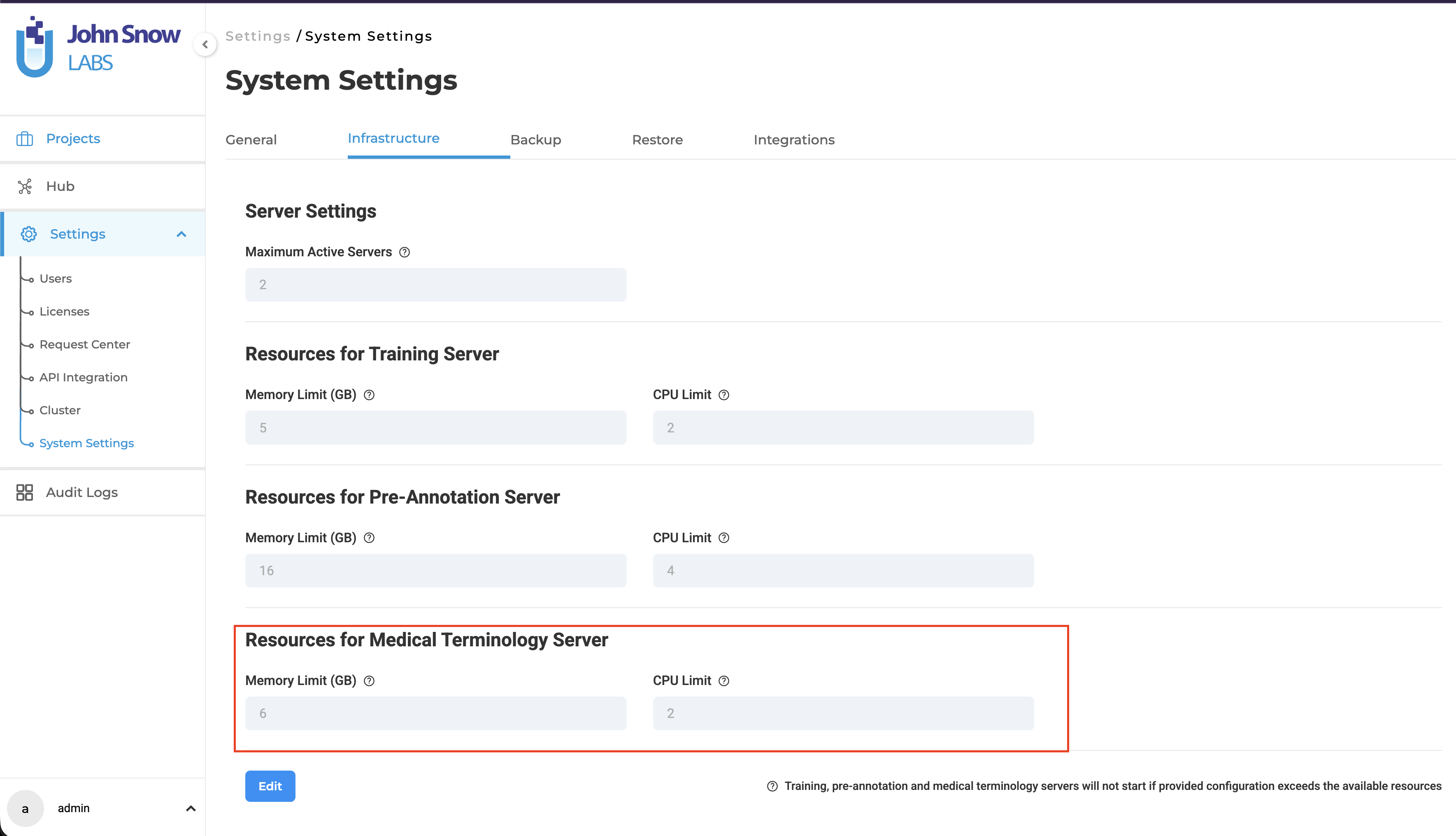
Default Resource Allocation:
- 2 CPU cores
- 6 GB memory
Teams can adjust these settings based on:
- Number of active projects using terminologies
- Size and complexity of terminology databases
- Expected query volume during annotation
- Infrastructure capacity and priorities
Cluster Management Visibility: The Cluster Management page displays terminology server status, including:
- Which terminologies are deployed on which server
- Server uptime and health status
- CPU and memory usage
- Active projects using each server
This visibility gives administrators clear insight into terminology infrastructure without requiring manual coordination across projects.
Licensing and Credit Consumption
Medical Terminology usage follows a clear and predictable licensing model aligned with the platform’s Healthcare pipeline approach.
Credit Consumption Rules:
Standalone Terminology Deployments:
- Consume 2 floating credits
- Apply when a Medical Terminology Server is deployed without an associated NER pipeline
Parallel Deployments with NER Pipelines:
- Consume 0 additional credits
- Apply when a Medical Terminology Server is deployed alongside an NER pre-annotation pipeline within the same project
- The terminology deployment runs in parallel with the NER pipeline without additional credit consumption
License Requirements:
- Downloading Medical Terminologies requires a valid license
- Healthcare scope license is required for healthcare-specific terminologies (ICD-10, LOINC, CPT, SNOMED CT, RxNorm)
- MeSH terminology follows the same licensing model
This licensing approach makes it economical to combine entity extraction and code resolution in the same project while maintaining flexibility for terminology-only workflows.
Benefits of Medical Terminologies
Clinical Standardization: Apply recognized healthcare coding systems directly within annotation workflows for consistent structured outputs. Annotations carry both surface forms and standardized codes, enabling interoperability with downstream healthcare systems.
Improved Annotation Accuracy: Reduce coding inconsistency by enabling automated resolution and guided manual lookup. Annotators can verify both entity labels and medical codes in a single pass, improving overall annotation quality.
Reusable Shared Assets: Download terminology resources once and reuse them across multiple projects. Teams working on different annotation initiatives can draw from the same terminology without redundant setup or licensing.
Efficient Infrastructure Usage: Consolidate multiple terminologies into a single server and reuse existing active deployments where possible. The platform’s intelligent server management reduces infrastructure overhead while maintaining functionality.
Predictable Credit Consumption: Clear distinction between standalone terminology deployments (2 credits) and zero-cost parallel deployments with NER pipelines. Teams can plan resource allocation with full visibility into credit usage.
Better Interoperability: Generate outputs aligned with standardized clinical coding systems for downstream healthcare analytics, billing systems, research databases, and regulatory submissions. Bridging the gap between extracted entities and coded data happens within the annotation workflow.
Example Workflow
A healthcare annotation team is building a project to label diagnoses and laboratory findings from patient records:
- Download Terminologies:
- Navigate to the Hub
- Download ICD-10 and LOINC terminology assets
- Configure Project:
- Create a new NER project
- Go to Configuration → Reuse Resource → Customize Label
- Link ICD-10 to diagnosis labels
- Link LOINC to lab-related labels
- Save configuration
- Deploy Servers:
- Navigate to the Task page
- Click Pre-Annotate → Create Server → Deploy
- Both Medical Terminology Server and Pre-Annotation Server are deployed together
- Automated Resolution:
- Pre-annotation runs on imported documents
- Entities are extracted and medical codes are automatically resolved
- Annotators see entity spans with attached ICD-10/LOINC codes
- Manual Lookup During Review:
- Annotators review automated results
- When needed, use manual terminology lookup to verify or correct codes
- Select alternative codes interactively within the annotation interface
- Server Reuse:
- Create a second project requiring the same terminologies
- The system automatically reuses the existing Medical Terminology Server
- No additional deployment or credit consumption needed
- Zero-Cost Parallel Deployment:
- Because the project includes both an NER pipeline and Medical Terminologies
- The terminology deployment runs alongside the NER pipeline
- No additional floating credits are consumed
This workflow improves consistency, reduces manual effort, and ensures clinical annotations remain aligned with standardized healthcare terminologies throughout the entire annotation lifecycle.