Generative AI Lab offers tight integration with NLP Models. Any compatible model and embeddings can be downloaded and made available to the Generative AI Lab users for pre-annotations either from within the application or via manual upload.
NLP Models HUB page is accessible from the left navigation panel by users in the Admins group.
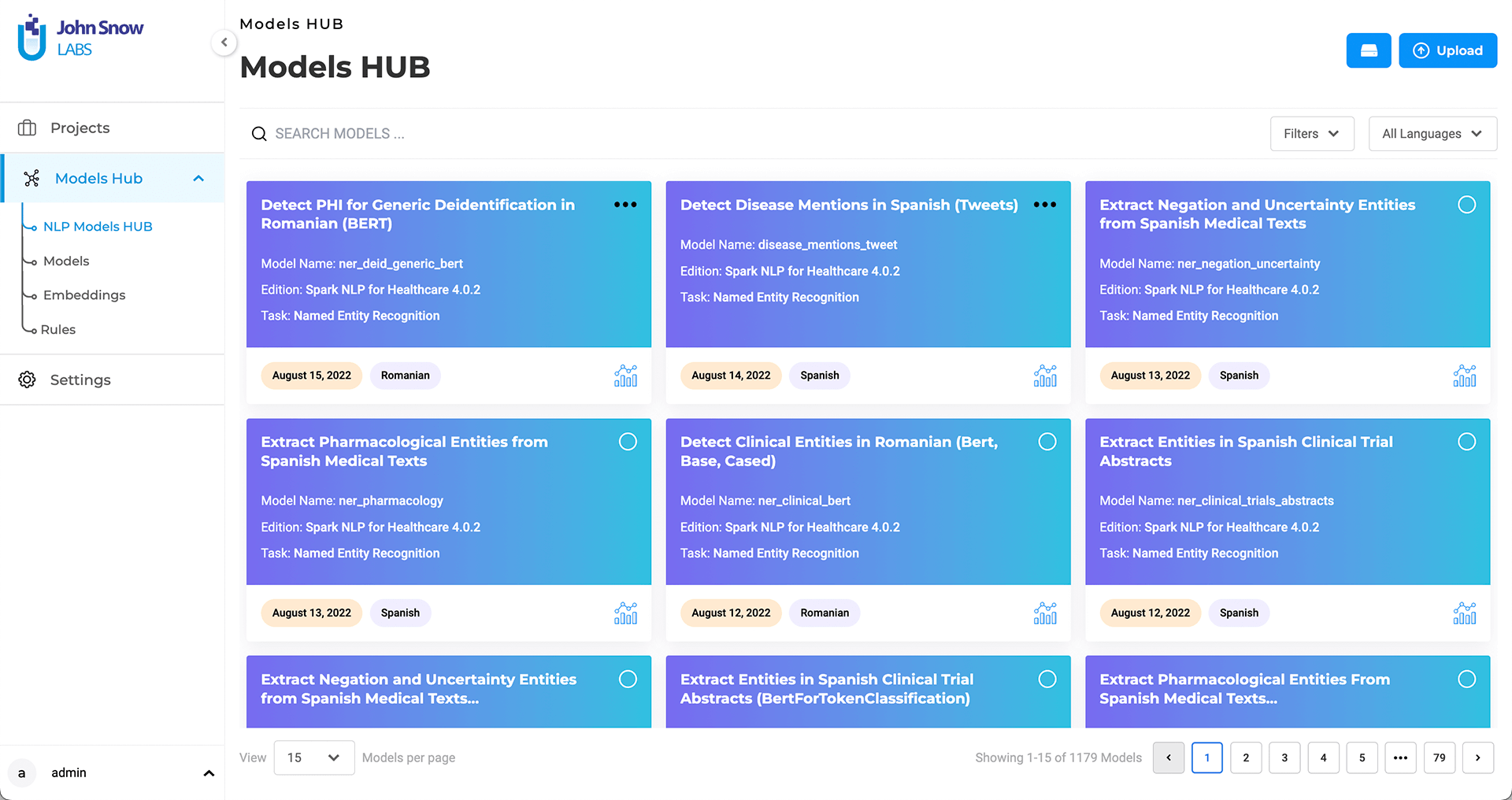
The Models Hub page lists all the pre-trained models and embeddings available from the NLP Models Hub that are compatible with the Lab’s current Spark NLP version.
Generative AI Lab automatically detects the active Spark NLP library version and filters available models to ensure compatibility.
Search
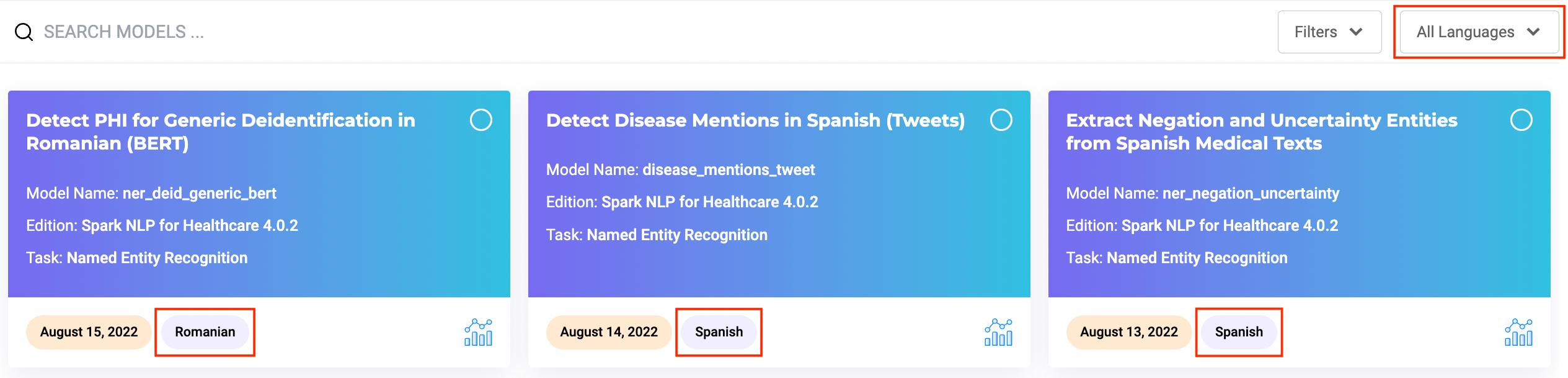
Search features are offered to help users identify the models they need based on their names. Additional information such as Library Edition, task for which the model was build as well as publication date are also available on the model tile.
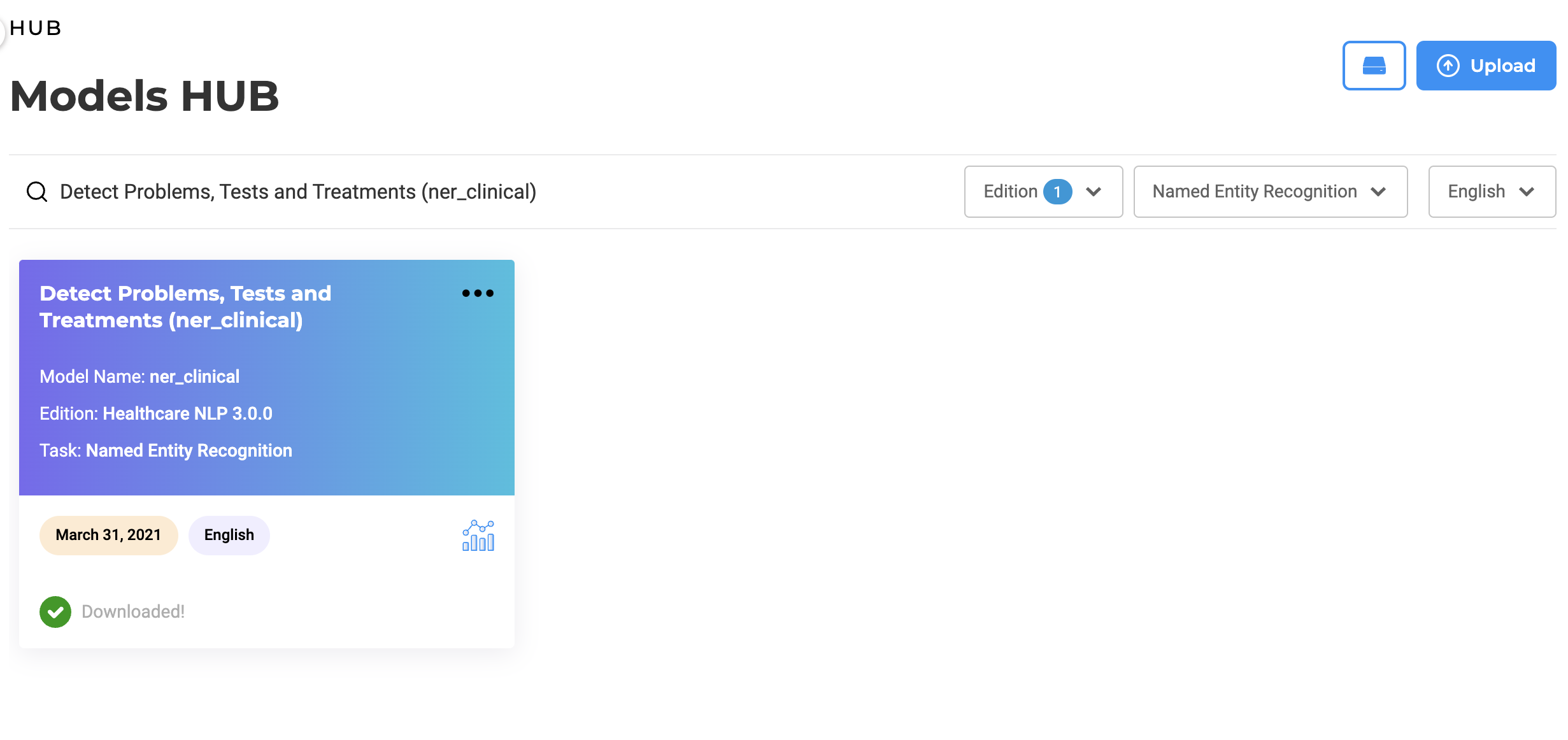
Language of the model/embeddings is also available as well as a direct link to the model description page on the NLP Models Hub where you can get more details about the model and usage examples.
Filter
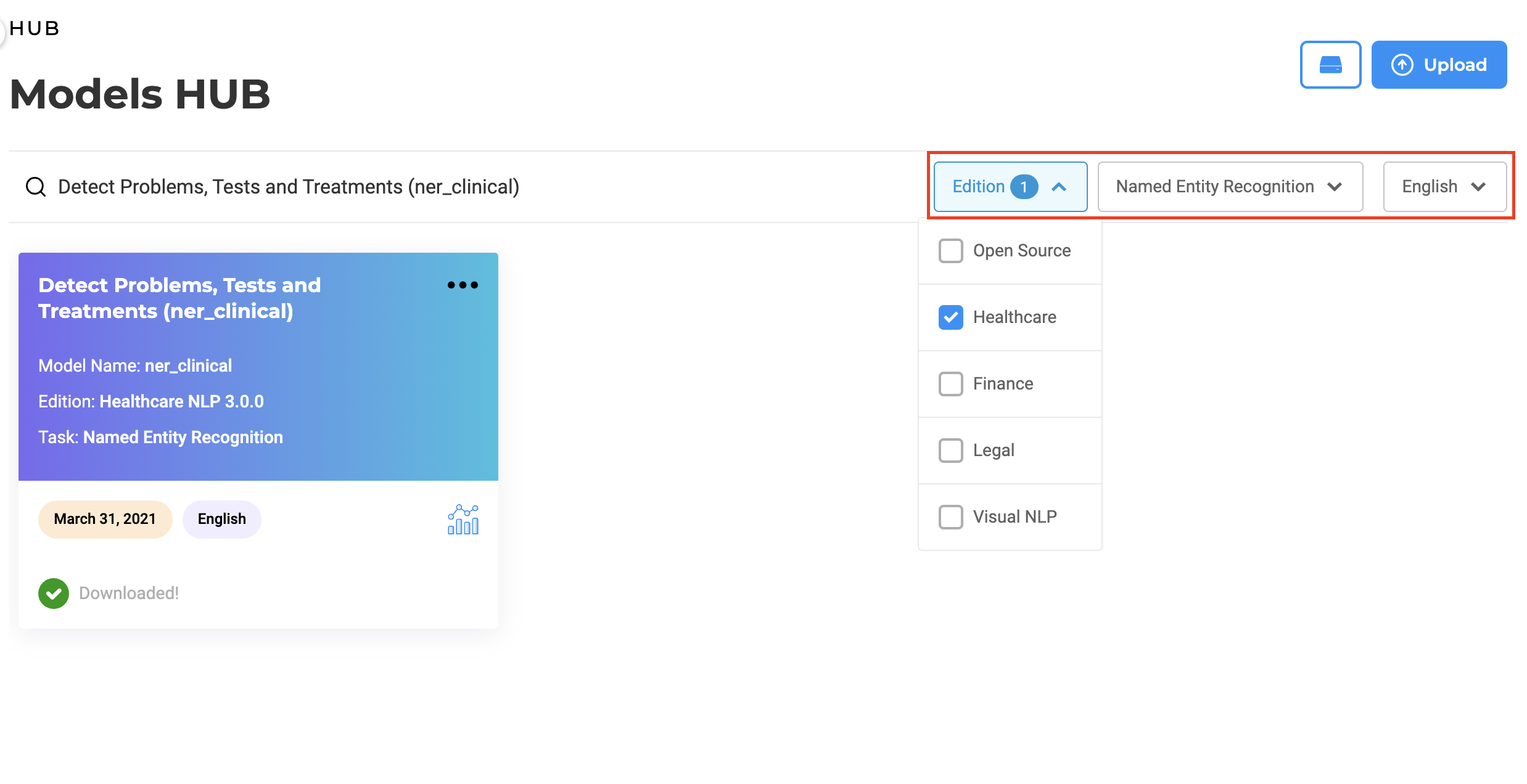
Users can use the Edition filter to search models specific to an edition. It includes all supported NLP editions: Healthcare, Opensource, Legal, Finance, and Visual. When selecting one option, e.g. “Legal”, users will be presented with all available models for that specific domain. This will ease the exploration of available models, which can then easily be downloaded and used within Generative AI Lab projects.
To make searching models/embeddings more efficient, Generative AI Lab offers a Language filter. Users can select models/embeddings on the Models Hub page according to their language preference.
Download
By selecting one or multiple models from the list, users can download those to the Generative AI Lab. The licensed (Healthcare, Visual, Finance or Legal) models and embeddings are available to download only when a valid license is present.
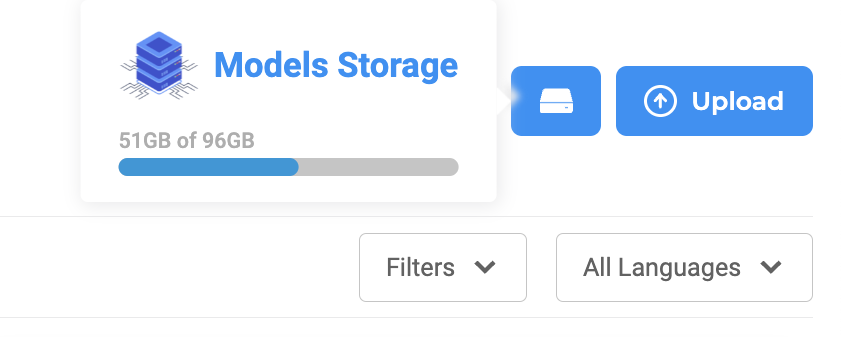
One restriction on models download/upload is related to the available disk space. Any model download requires that the double of its size is available on the local storage. If enough space is not available then the download cannot proceed.
Disk usage view, search, and filter features are available on the upper section of the Models Hub page.
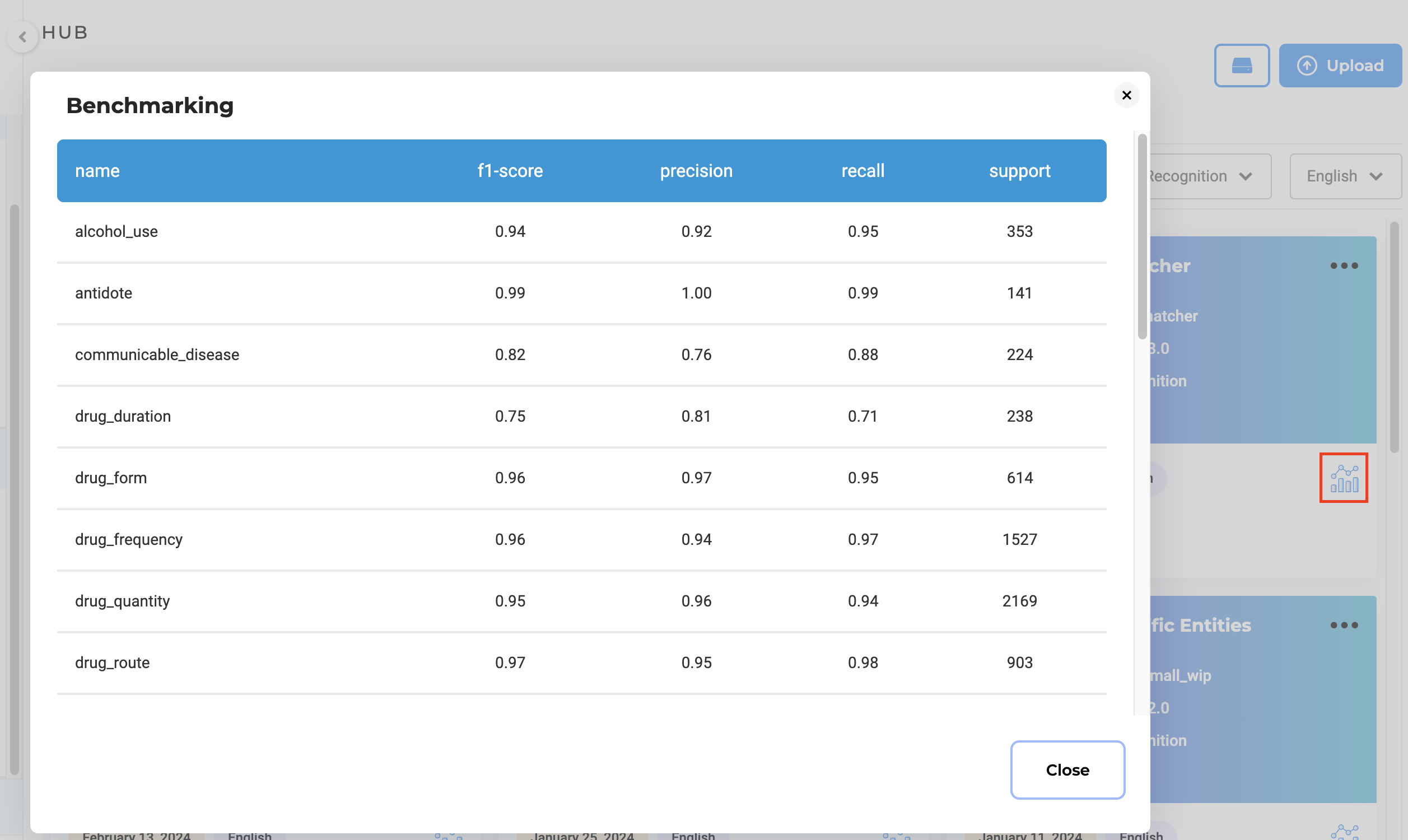
Benchmarking
For the licensed models, benchmarking information is available on the Models Hub page. To check this click on the icon on the lower right side of the model tile. The benchmarking information can be used to guide the selection of the model you include in your project configuration.